LH - Tutorials
Brains, Minds, and Machines Summer Course Tutorials
The annual Brains, Minds and Machines Summer Course includes many tutorials on key computational and empirical methods used in research on intelligence. Lecture videos and slides are provided for each tutorial, and some also include resources to support hands-on computer activities. The summer course is aimed at graduate students and postdocs, but many of the tutorials are accessible to undergraduates with basic math background (e.g. calculus, linear algebra, probability and statistics).
Andrzej Banburski, MIT
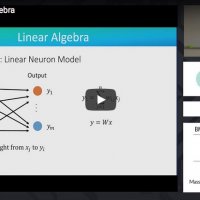
Introduction to concepts from linear algebra needed to understand Principal Components Analysis (PCA): vectors, matrices, matrix multiplication and other operations, data transformations, solving linear equations, and PCA.
Andrei Barbu, MIT
Introduction to probability, covering uncertainty, simple statistics, random variables, independence, joint and conditional probabilities, probability distributions, Bayesian analysis, graphical models, mixture models, and Hidden and Markov models.
Ethan Meyers, Hampshire College/MIT
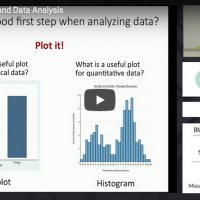
Overview of statistics including descriptive statistics, data plots, sampling distributions, statistical inference, estimation, regression, confidence intervals, and hypothesis testing.
Kevin Smith, MIT
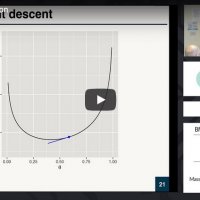
Introduction to optimization focused on basic concepts, terminology, and its applications. Topics include likelihood and cost functions, single and multi-variable optimization, and optimization for machine learning (stochastic gradient descent, regularization, sparse coding, and momentum).
Lorenzo Rosasco, Universita' di Genova/Istituto Italiano di Tecnologia/MIT
Overview of basic concepts and theory underlying many common machine learning methods and their application to different kinds of problems. This theory is complemented by computer labs in MATLAB that explore the behavior of machine learning methods in practice.
Eugenio Piasini, UPenn & Yen-Ling Kuo, MIT
Overview of supervised learning with neural networks, convolutional neural networks for object recognition, and recurrent neural networks, with a brief introduction to other deep learning models such as auto-encoders and generative models.
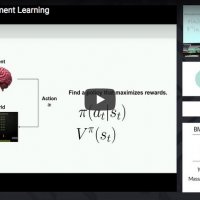
Xavier Boix & Yen-Ling Kuo, MIT
Introduction to reinforcement learning, its relation to supervised learning, and value-, policy-, and model-based reinforcement learning methods. Hands-on exploration of the Deep Q-Network and its application to learning the game of Pong.
Programming problems in computational neuroscience by Emily Mackevicius, on feedforward networks, estimating the voltage response of a neuron, edge detection in images, analyzing premotor neural data, recognizing facial expressions, and spike-triggered averaging.
Francisco Flores, MIT
Overview of basic neuroscience concepts, including the structure of neurons, membrane biophysics and the generation of action potentials, synapses, neural networks, sensory pathways in the brain, the motor system and sensory-motor integration.
Diego Mendoza-Halliday, MIT
Introduction to methods for recording, analyzing, and visualizing neural signals, focused on electrophysiology methods such as the patch clamp technique, intracellular and extracellular electrode recordings, local field potentials, ECoG, and EEG.
Frederico Azevedo, MIT
Introduction to ventral and dorsal pathways in the brain, hierarchy of the visual system and face processing areas, action pathways and decoding movement, selective visual attention, long and short-term memory, learning and synaptic plasticity.
Pouya Bashivan, MIT
Overview of the visual pathways from the retina to higher cortical areas; approaches to modeling neural behavior including spike triggered averaging, linear-nonlinear methods, generalized linear models, Gabor filters and wavelet transforms, and convolutional neural network models; and using models for neural prediction and control.
Kohitij Kar, MIT
Models for encoding and decoding visual information from neural signals in the ventral pathway that underlie core object and face recognition, testing neural population decoding models to explain human and monkey behavioral data on rapid recognition tasks.
Ethan Meyers, Hampshire College/MIT
Introduction to neural decoding methods to study the neural representations of sensory information in the brain to support recognition, their modulation by task-relevant information from top-down attention, and existence of sparse, dynamic population codes.
Mengmi Zhang, Harvard
Introduction to deep convolutional generative adversarial networks and their application to image reconstruction from brain signals. This technology may lead to more effective brain-computer interfaces.
Andrei Barbu, MIT
Overview of some challenges still facing the design of general computer vision systems, traditional image processing operations and applications of machine learning to visual processing.
Kohitij Kar, MIT
Introduction to common psychophysical methods, including magnitude estimation, matching, detection and discrimination, the two-alternative forced choice paradigm, psychometric curves and signal detection theory, and using Amazon Mechanical Turk for large-scale experiments.
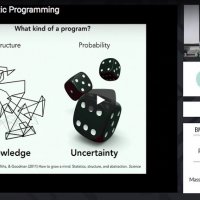
Kevin Smith, MIT
Probabilistic programming languages facilitate the implementation of generative models of the physical and social worlds that enable probabilistic inference about objects, agents, and events. This tutorial introduces WebPPL through example models and inference techniques.
Tomer Ullman, MIT/Harvard
Online notes on the Church language, introducing its syntax, basic primitives, and how to define functions that implement simple probabilistic models and inference methods. Concepts are explored through coding examples and exercises.
fMRI Methods and Analysis
Rebecca Saxe, MIT
This nine-part video series introduces the basics of fMRI and delves into many current methods for analyzing fMRI data, including univariate and multivariate analyses, multivoxel pattern analysis (MVPA), and representational similarity analysis.
AFNI is a set of C programs for processing, analyzing, and displaying anatomical and functional MRI data. This series of video tutorials introduces AFNI and is aimed at researchers who are already using fMRI in their work.
Nikos K. Logothetis, Max Planck Institute for Biological Cybernetics in Tübingen
Overview of technical advances that enable the integration of direct electrical stimulation of neural tissue with fMRI recordings, and application of this technology to study monosynaptic neural connectivity, network plasticity, and cortico-thalamo-cortical loops in the primate brain.
Nikos K. Logothetis, Max Planck Institute for Biological Cybernetics in Tübingen
Overview of the integration of concurrent physiological multi-site recordings with fMRI imaging, and it application to the study of dynamic connectivity related to system and synaptic memory consolidation in primates.
BCS Computational Tutorials
Seminar series led by graduate students and postdocs in the MIT Department of Brain and Cognitive Sciences (BCS), featuring tutorials on computational topics relevant to research on intelligence in neuroscience, cognitive science, and AI. Live tutorials include a lecture followed by “office hours” to work through problems and discuss how the material may be applied to participants’ research. Resources posted here include lecture videos created by CBMM, lecture slides, code and datasets for exercises, background references, and other supplementary material. These tutorials are aimed at graduate students and postdocs with some computational background, but who are not experts on these topics. This series was organized by Emily Mackevicius and Jenelle Feather, with financial support from the MIT Department of Brain and Cognitive Sciences.
Mehrdad Jazayeri & Josh Tenenbaum, MIT (2015)
An introduction to Bayesian estimation and its application to estimating visual contrast from neural activity.
Luke Hewitt, MIT (2018)
Overview of approximate inference methods for implementing Bayesian models, such as sampling-based methods and variational inference, and probabilistic programming languages for implementing these models.
Sam Gershman, Harvard (2017)
This tutorial introduces the basic concepts of reinforcement learning and how they have been applied in psychology and neuroscience. Hands-on exercises explore how simple algorithms can explain aspects of animal learning and the firing of dopamine neurons.
Phillip Isola, MIT (2015)
Introduction to using and understanding deep neural networks, their application to visual object recognition, and software tools for building deep neural network models.
Larry Abbott, Columbia (2015)
Introduction to recurrent neural networks and their application to modeling and understanding real neural circuits.
Emily Mackevicius & Greg Ciccarelli, MIT (2016)
Introduction to dimensionality reduction and the methods of Principal Components Analysis and Singular Value Decomposition.
Sam Norman-Haignere, MIT (2016)
Dimensionality Reduction using the method of Independent Components Analysis, and its application to the analysis of fMRI data.
Alex Williams, Stanford (2017)
This tutorial covers matrix and tensor factorizations, a class of dimensionality-reduction methods that includes PCA, independent components analysis (ICA), and non-negative matrix factorization (NMF).
Harini Suresh & Nick Locascio, MIT (2017)
Introduction to Long Short Term Memory networks (LSTMs), a type of recurrent neural network that captures long term dependencies, often used for natural language modeling and speech recognition, with an introduction to TensorFlow.
Anima Anandkumar, UC Irvine (2016)
Guaranteed Machine Learning using tensor methods, with examples of learning probabilistic models and representations.
Ethan Meyers, Hampshire College & MIT (2015)
Introduction to the Neural Decoding Toolbox, used to analyze neural data from single cell recordings, fMRI, MEG, and EEG, to understand the information contained in this data and how it is encoded and decoded by the brain.
Seth Egger, MIT (2016)
Introduction to linear dynamical systems, stochastics, and the evolution of probability density, with application to modeling the generation of neural signals.
Emily Mackevicius & Andrew Bahle, MIT (2018)
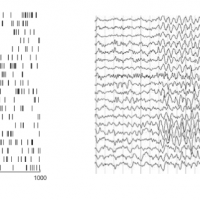
Introduction to the SeqNMF tool for unsupervised discovery of temporal sequences in high-dimensional datasets, applied to neural and behavioral datasets.
Mark Goldman, UC Davis (2016)
This tutorial describes how to apply linear network theory to the analysis of neural data. It introduces the concept of “sloppy models,” in which model parameters are poorly constrained by available data.
Andre M. Bastos, MIT (2016)
Review of methods used in invasive and non-invasive electrophysiology to study the dynamic connections between neuronal populations, including common pitfalls that arise in functional connectivity analysis and how to address these problems.
Jai Bhagat & Caroline Moore-Kochlacs, MIT (2017)
This tutorial introduces common electrophysiology setups and describes the spike sorting workflow, from filtering the raw voltage traces to assessing the quality of the final spike clusters, ending with spike-sorting simulations in MATLAB.
Pengcheng Zhou, Columbia (2017)
Reviews existing methods to extract neurons from raw calcium imaging video data and then focuses on how to use the CNMF-E method to analyze 1-photon microendoscopic data in MATLAB.
Eric Denovellis, BU (2017)
This tutorial focuses on best coding practices to develop code that is reusable, sharable, and bug free. It highlights issues such as documentation, version control, and unit testing, through hands-on computer exercises.